Membrane proteins make up an estimated 30% of the proteome and 50% of drug targets, but less than 2% of the structures in the protein data bank. This dearth of structural information is primarily due to the experimental difficulty of inducing membrane proteins to crystallize or reconstituting them in solubilizing media. Solid-state NMR is an emerging technique that is increasingly used to investigate membrane proteins under close to native conditions. Membrane samples can be macroscopically aligned with the membrane normal along a specific direction, often parallel or perpendicular to the magnetic field. This is exploited in both static experiments on membrane samples and in the solution-state residual dipolar coupling (RDC) experiment, which has proven extremely useful for obtaining long-range distance constraints, but is limited to the case of very weak alignment. In solid-state NMR, anisotropic interactions are typically removed via magic angle spinning (MAS). Heteronuclear or homonuclear dipolar couplings are often selectively reintroduced using pulse sequences, because they depend on internuclear distances. The standard recoupling sequences are difficult to implement in mobile samples, where the dipolar interactions are partially averaged by rotational diffusion; optimized sequences are needed.
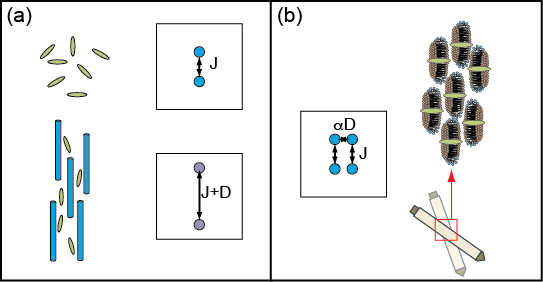
In switched-angle spinning (SAS) NMR, the spinning axis is varied in two correlated Fourier dimensions, allowing isotropic spectra and anisotropic couplings to be collected in the same experiment. SAS measures the isotropic spectrum needed to differentiate chemical sites, while providing valuable information that would be lost in a simple MAS experiment, all without changing the any concentrations or the temperature (shown schematically below). This approach makes the mobility of the sample an asset instead of a liability. Preliminary SAS and VAS experiments have been performed in membrane mimetics, illustrating the potential of the technique, however technical limitations have prevented their widespread application.
Obtaining high-resolution SAS spectra of membrane proteins in their native environment requires a major research effort directed toward building new SAS probes with the high radio frequency (rf) homogeneity, tuning stability, and temperature control necessary for long multidimensional experiments on biological samples. These requirements contrast with the design parameters for previous hardware for quadrupolar materials and other rigid solids, in which the major criteria included high-power rf irradiation and fast spinning. The biggest challenge in SAS probe design is the need to maintain a stable electrical contact between the coil, which is rapidly reorienting during the experiment and the rest of the probe circuit, which is static. For experiments on oriented membrane samples, the filling factor must be maximized to use the small samples available, and the connection points must be robust enough to withstand long experiments with many points in the indirect dimension. The increased stringency of these requirements is somewhat compensated by the decreased necessity for high excitation or decoupling fields. The direction of the rf field should remain perpendicular to B0 regardless of the spinning angle. The switching speed must be much faster than the 1H longitudinal relaxation time T1 for a typical hydrated solid protein sample (about 1s). Two prototype probes that demonstrate fulfillment of these criteria have been constructed in the Martin laboratory. Pneumatic switching results in a minimum switching time of 17 ms, with stable spinning at 5 kHz, which is sufficient for membrane samples. We have used variations of two transverse coil designs, the slotted tube resonator and the saddle coil, and found them to have complementary features. We have also investigated the dynamics of cross-polarization, an important building block of solid-state NMR experiments in which magnetization is transferred from abundant, high gyromagnetic-ratio nuclei to rare, low gyromagnetic-ratio nuclei, as a function of spinning angle. Although the essential phenomena involved have been described in the early literature on MAS and cross-polarization, collecting the full data set from 0° to 90° is experimentally challenging because it requires a probe where the rf fields applied to both 1H and 13C are independent of the spinning angle. Currently, we are constructing three-channel 1H/13C/15N probes to perform 3D experiments for resonance assignment of proteins. We are also investigating the performance of homonuclear and heteronuclear decoupling sequences, which also strongly depend on the spinning angle, and designing symmetry-based sequences for decoupling. Finally, there are many interesting applications to investigate, first and foremost membrane-associated proteins.
